Genome AssemblyHuman Whole Genome ResequencingFull-length Transcriptome SequencingMicrobial MetagenomicsWhole-Plasmid Sequencing and Assembly
 De novo sequencing of the genome of a species with no reference sequence information. Genome mapping of the species is obtained through assembly and linking assisted by bioinformatic analysis methods. The whole-genome mapping can be used to construct the genomic database of that species, which provides DNA sequence information for further genome mining and functional validation.
De novo sequencing of the genome of a species with no reference sequence information. Genome mapping of the species is obtained through assembly and linking assisted by bioinformatic analysis methods. The whole-genome mapping can be used to construct the genomic database of that species, which provides DNA sequence information for further genome mining and functional validation. Human pangenome research
Animal genome assembly
Plant genome assembly
Bacteria genome assembly
Technical Advantages
CycloneSEQ achieves uniform coverage with long read lengths and enables the resolution of complex genomes.
No GC bias, allowing for accurate sequencing of GC-rich regions.Long read lengths that ensure uniform coverage of highly repetitive regions.
CycloneSEQ's long-read data provides contig connectivity, thereby improving the continuity of genome assembly.
Delivers reads longer than 100 kb, with the maximum read length reaching the Mbp level.Can be used for haplotype analysis.
The extended read lengths provided by CycloneSEQ aid in the efficient assembly of finished microbial genomes.
Easy 0-gap and N-free genome assembly.Increased copy number and improved completeness of 16S rRNA and other genes.
Application Case
CycloneSEQ data used for the T2T genome assembly of Oryza longistaminata Asian cultivated rice (Oryza sativa) is one of the world's most important food crops, and its wild relatives serve as an invaluable genetic foundation for rice breeding. Compared with Asian cultivated rice, African wild rice (Oryza longistaminata) has advantages in traits such as resistance to increased biomass production, rhizome-based vegetative propagation, and resistance to biotic stresses. However, previous reference genomes of African wild rice contained numerous gaps and incomplete regions, limiting detailed studies on it. Based on the CycloneSEQ nanopore sequencing platform, a 343 Mb telomere-to-telomere (T2T) genome assembly of African wild rice (O. longistaminata) was completed, providing a valuable resource for the exploration and utilization of beneficial alleles in wild relatives of Asian cultivated rice. Source: Telomere-to-telomere African wild rice (Oryza longistaminata) reference genome reveals segmental and structural variation bioRxiv 2024.09.05.611405; doi: https://doi.org/10.1101/2024.09.05.611405
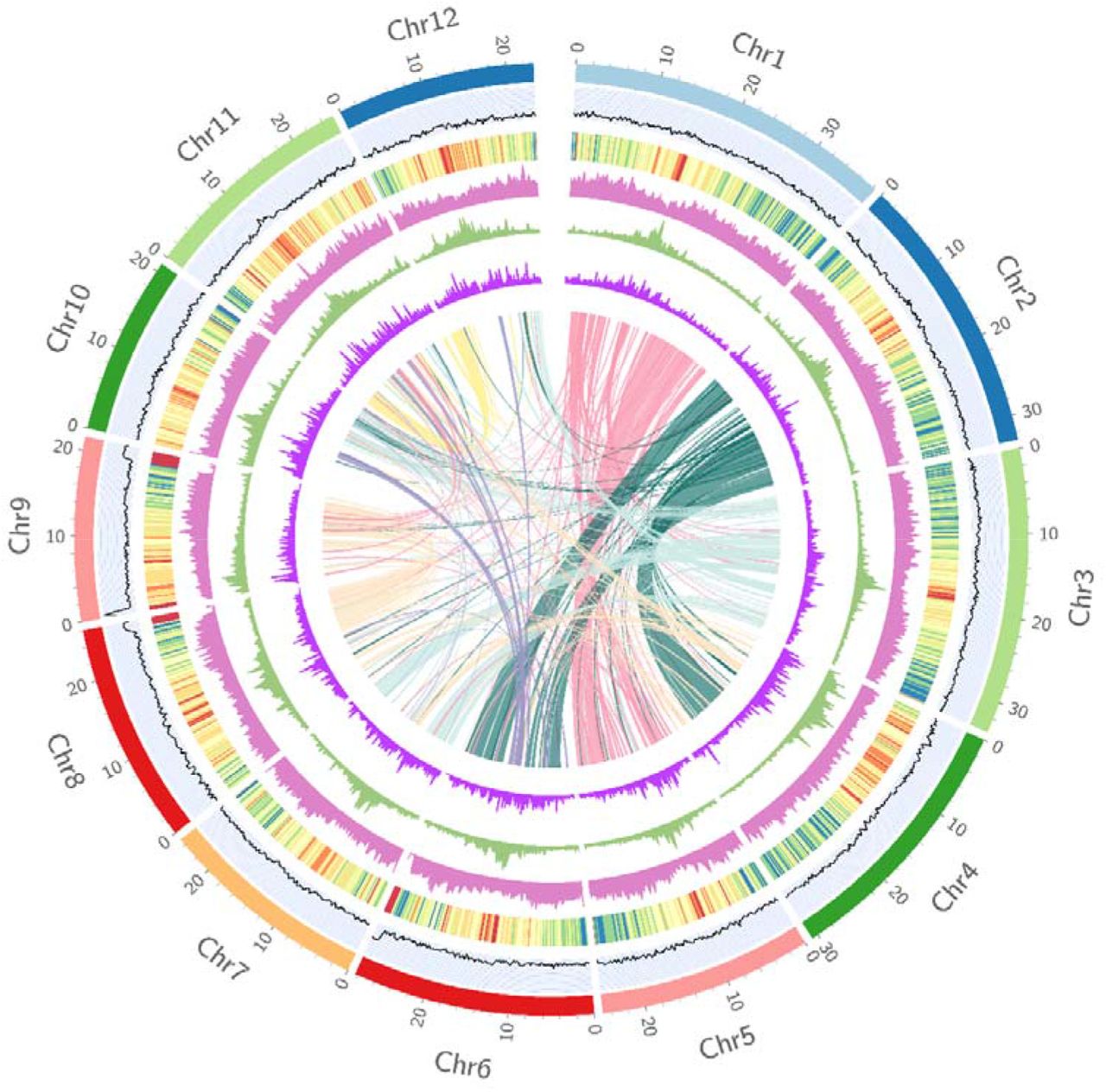 Figure 1 CycloneSEQ data used for the T2T genome assembly of Oryza longistaminata
Figure 1 CycloneSEQ data used for the T2T genome assembly of Oryza longistaminata The bearded dragon (Pogona vitticeps) of the family Agamidae is one of the most popular pet reptiles in the world. Its ability to be bred in captivity has also made it a model organism, particularly for studies on the origin of sex chromosomes and the mechanism of sex determination. Result: A chromosome-level reference genome for the bearded dragon was established using whole-genome and long-read sequencing of a captive-bred ZZ male individual, employing BGI's CycloneSEQ nanopore sequencing and MGI's DNBSEQ gene sequencing technologies. The new reference genome is approximately 1.8 Gb in length, with an N50 length of 202.5 Mb, and all assembled fragments are anchored onto 16 chromosomes. Source: A near-complete genome assembly of the bearded dragon Pogona vitticeps provides insights into the origin of Pogona sex chromosomes bioRxiv 2024.09.05.611321; doi: https://doi.org/10.1101/2024.09.05.611321
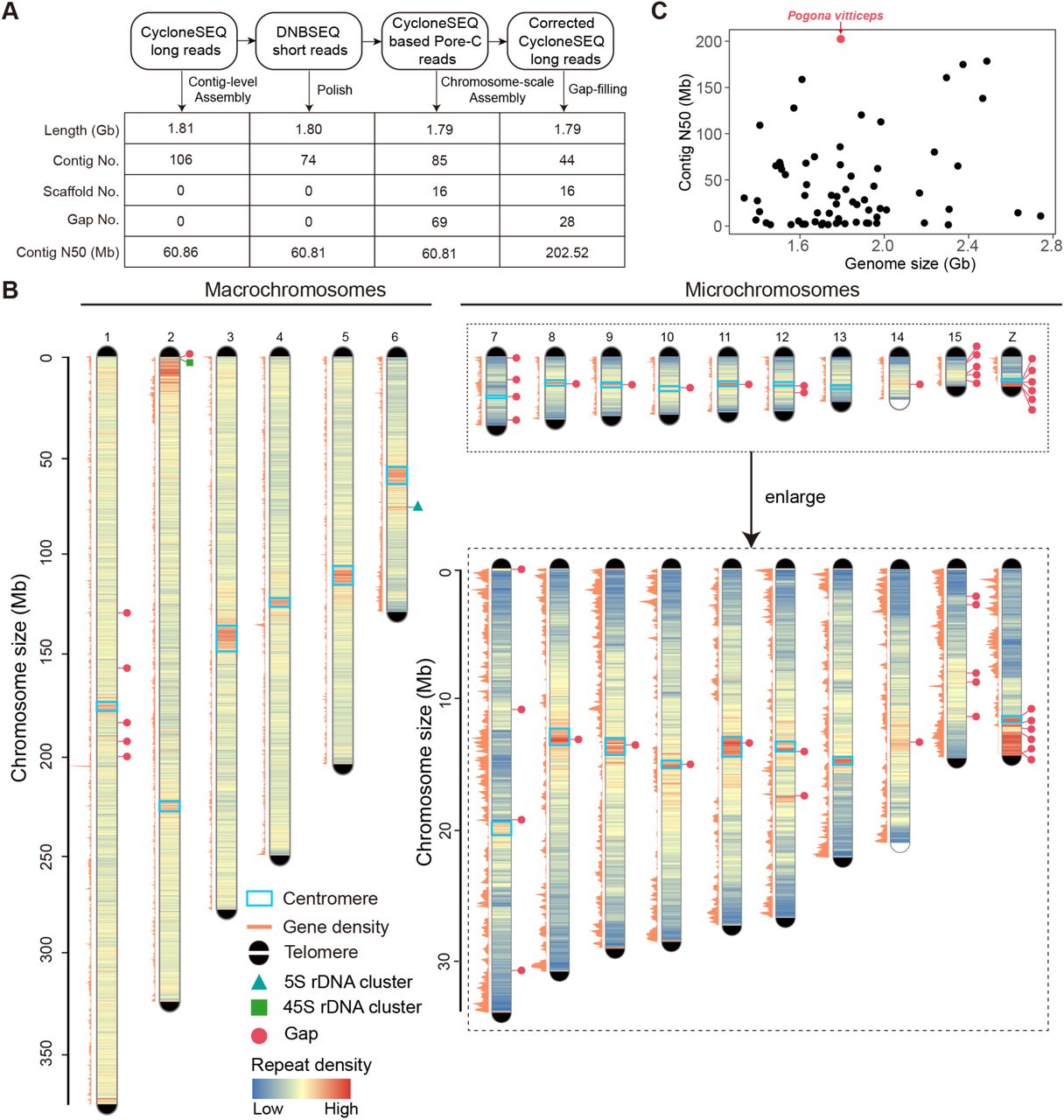 Figure 2 Near-complete genome assembly of the bearded dragon Pogona vitticeps
Figure 2 Near-complete genome assembly of the bearded dragon Pogona vitticeps

